Introduction¶
Prerequisites¶
Matlab / Octave Advanced beginner level.
fMRI analysis :ref:`advanced beginner level <cogneuro_experience>.
Goals of this course¶
Describe the typical MVPA approaches (correlation analysis, classification analysis, representational similarity analysis applied to both regions of interest and across the whole brain in a searchlight approach) described in the literature.
- Learn how to use CoSMoMVPA to perform these analyses:
Understand the dataset structure to represent both the data itself (e.g. raw measurements or summary statistics) and its attributes (e.g. labels of conditions (targets), data acquisition run (chunks).
See how parts of the data can be selected using slicing and splitting, and combined using stacking
Introduce measures that compute summaries of the data (such as correlation differences, classification accuracies, similarity to an a prior defined representational similarity matrix) that can be applied to both a single ROI or in a searchlight.
Make yourself an independent user, so that you can apply the techniques learnt here to your own datasets.
Not covered in this course¶
Preprocessing of fMRI data
Learning to use Matlab / Octave
Other dataset types than volumetric fMRI data (MEEG, surface-based fMRI)
How to become a CoSMoMVPA developer
Code and data needed for this workshop¶
Sample Dataset¶
The dataset used here contains preprocessed data for 8 subjects from [CGG+12]. In this experiment, participants were presented with categories of six animals: 2 primates: monkeys and lemurs; 2 birds: mallard ducks and yellow-throated warblers; and 2 bugs: ladybugs and luna moths.
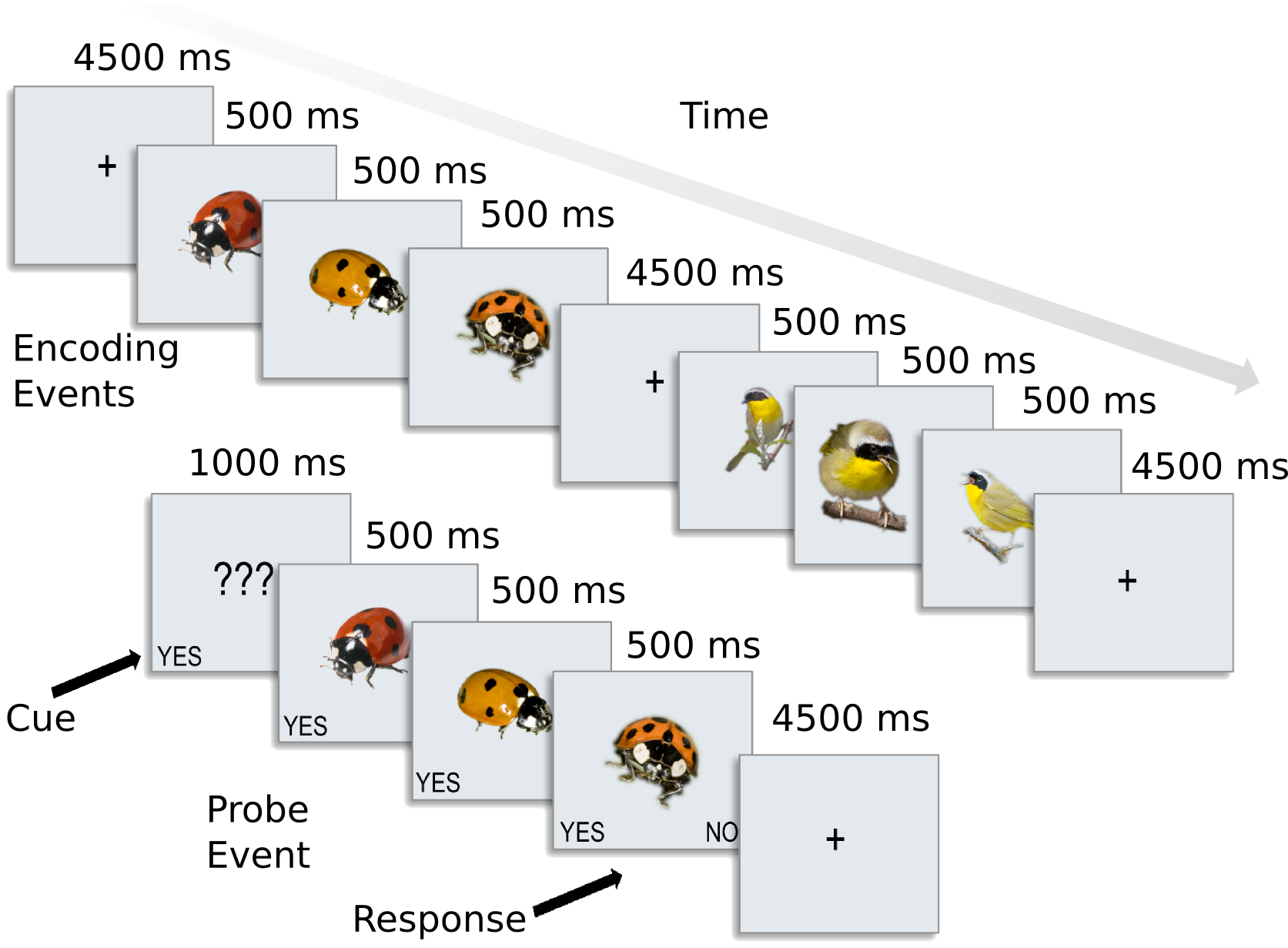
For each participant, the following data is present in the ak6 (for Animal Kingdom, 6 species) directory:
- s0[1-8]/ This directory contains fMRI data from 8 of the 12
participants studied in the experiment reported in
Connolly et al. 2012 (Code-named 'AK6' for animal
kingdom, 6-species). Each subject's subdirectory
contains the following data:
- glm_T_stats_perrun.nii A 60-volume file of EPI-data preprocessed using
AFNI up to and including fitting a general linear
model using 3dDeconvolve. Each volume contains the
t-statistics for the estimated response to a one
of the 6 stimulus categories. These estimates were
calculated independently for each of the 10 runs
in the experiment.
- glm_T_stats_even.nii Data derived from glm_T_stats_perrun.nii.
- glm_T_stats_odd.nii Each is a 6-volume file with the T-values averaged
across even and odd runs for each category.
- brain.nii Skull-stripped T1-weighted anatomical brain image.
- brain_mask.nii Whole-brain mask in EPI-space/resolution.
- vt_mask.nii Bilateral ventral temporal cortex mask similar to
that used in Connolly et al. 2012.
- ev_mask.nii Bilateral early visual cortex mask.
Also present are model similarity structures, which you can see here:
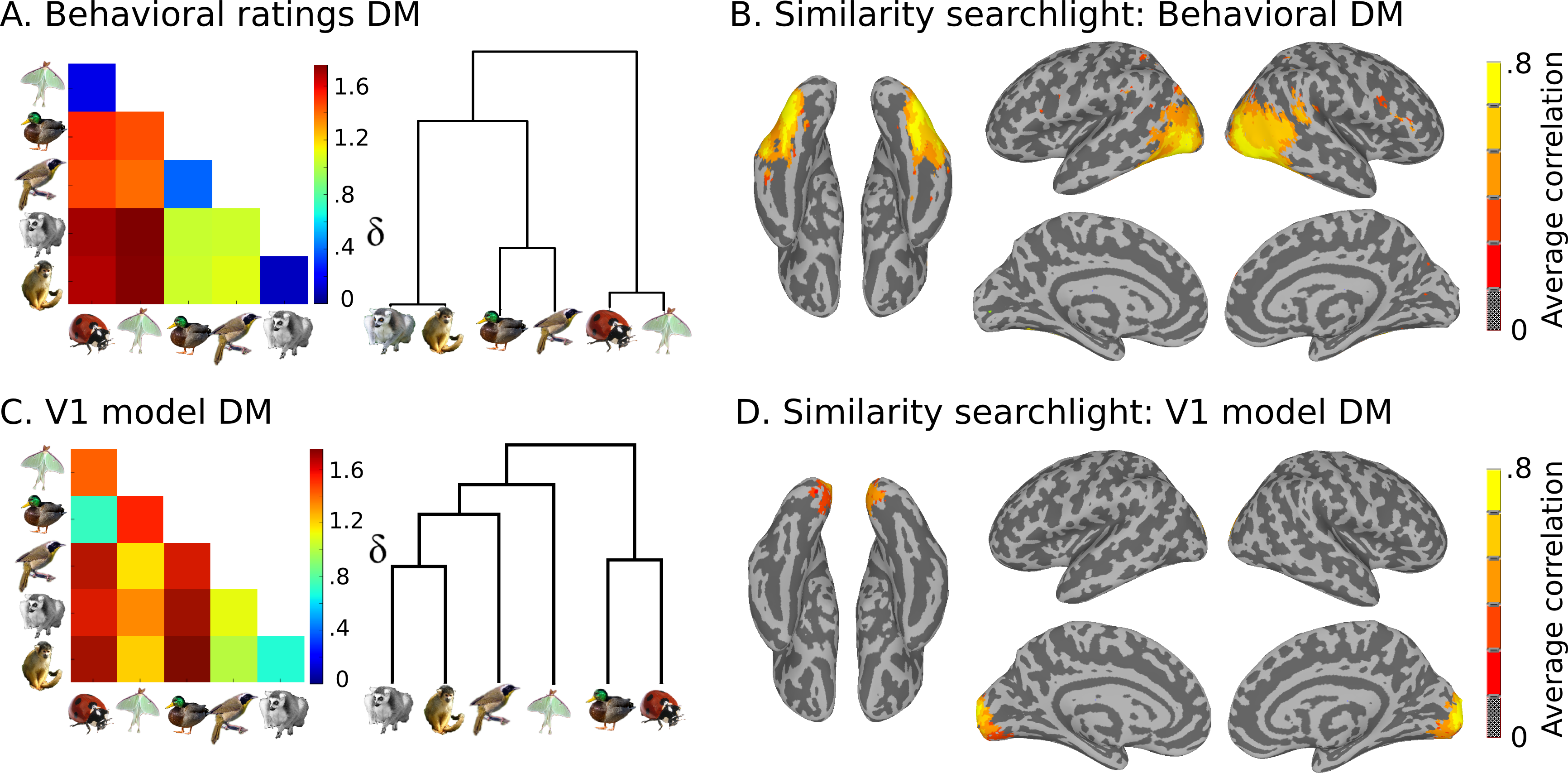
This data is stored in the models directory:
- models
- behav_sim.mat Matlab file with behavioural similarity ratings.
- v1_model.mat Matlab file with similarity values based on
low-level visual properties of the stimuli.
To cover in this course¶
CoSMoMVPA concepts and dataset structure.
Basic operations on datasets.
Introduction to common MVPA measures:
correlation difference
classification accuracy
representational similarity matching
Common MVPA techniques:
ROI analysis
Searchlight analysis

